Cutting-edge molecular biology techniques confirm the long-held but never proven assumption that toenail fungus and athlete’s foot can spread and that the fungus can infect family members living in the same household. Findings have important implications regarding the effective and timely treatment of these infections to prevent their spread throughout the family.
A collaboration between experts in dermatology, epidemiology, and mycology, the study was conducted at five centers led by Dr. Mahmoud Ghannoum, Professor of Dermatology at University Hospitals of Cleveland. The study was supported with funding from Novartis Pharmaceuticals.
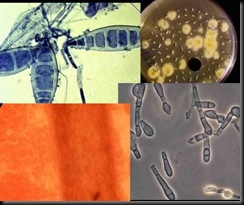
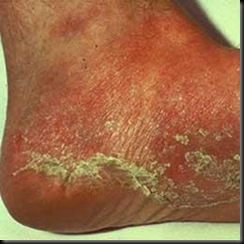
Affecting more than 35 million people, onychomycosis is a common, chronic, and highly resistant fungal infection in which affected nails become discolored, brittle, thickened, and flaky. Onychomycosis often results from tinea pedis, another common fungal skin infection affecting 10% of the general population at any time. Both infections are caused by tiny organisms called dermatophytes; the most common dermatophytes are Trichophyton rubrum (T. rubrum), Trichophyton mentagrophytes (T.mentagrophytes), and Epidermophyton floccosum (E. floccosum).
While it has long been assumed that dermatophytes spread from person to person, there has never been medical or scientific evidence supporting this contention. By providing scientific confirmation that dermatophytes can be transmitted among family members, this study suggests that toenail fungus and tinea pedis are infections that should be treated aggressively to help prevent their spread.
The prospective study enrolled 57 families in which at least one member, the “index person,” had either nail fungus and/or athlete’s foot. Of these, 19 families were identified in which at least two members were infected. Using conventional and culture growth methods to examine a total 110 toenail and skin samples taken from every participant, researchers determined that the vast majority of samples (80.4 %) were of the species T. rubrum, while most of the others were infected with T. mentagrophytes (14.3%), and E. floccosum. The identity of each species was confirmed using polymerase chain reaction (PCR), a molecular technique in which a species-specific fragment of DNA is amplified millions of times. Using a more sophisticated molecular biology technique called restriction fragment length polymorphism (RFLP), these amplified DNA molecules are further analyzed. Combining these two techniques, researchers were able to determine the identity of each strain based on unique DNA band patterns.
Results showed transmission of identical dermatophytes among 42% (8/19) of families. Among the families where spread of infection among members was detected, 87.5% (7/8) were infected with T. rubrum. Analysis of DNA band patterns using RFLP revealed that 62% (5/8) of the families were infected with the same strain of T. rubrum (Type D). Matching the DNA of dermatophytes among members of the same household clearly indicated that each family member had the identical strain of dermatophyte, and confirmed that the infection had been transmitted from one family member to the other either directly or indirectly.
In addition, the results of a questionnaire answered by the index person and family members identified the following as factors that significantly correlated with spread of infection in a household: past history of onychomycosis and tinea pedis, having tinea pedis for more than 10 years, and the occurrence of tinea pedis with scaling of the skin on the side of the foot.
The novel findings of this research suggest areas for future study, looking at factors that might determine why some family members became infected while others did not, as well as why the infection spread only within some of the families. Whether some strains may be more virulent, and if some individuals may be genetically susceptible to fungal infection, are other questions this study raises for future investigation
Source: American Society for Microbiology

No comments:
Post a Comment